| Issue |
A&A
Volume 618, October 2018
|
|
|---|---|---|
| Article Number | A13 | |
| Number of page(s) | 13 | |
| Section | Numerical methods and codes | |
| DOI | https://doi.org/10.1051/0004-6361/201732493 | |
| Published online | 05 October 2018 | |
Vaex: big data exploration in the era of Gaia⋆,⋆⋆
Kapteyn Astronomical Institute, University of Groningen, PO Box 800, 9700 AV Groningen, The Netherlands
e-mail: This email address is being protected from spambots. You need JavaScript enabled to view it.
Received:
19
December
2017
Accepted:
20
May
2018
Abstract
We present a new Python library, called vaex, intended to handle extremely large tabular datasets such as astronomical catalogues like the Gaia catalogue, N-body simulations, or other datasets which can be structured in rows and columns. Fast computations of statistics on regular N-dimensional grids allows analysis and visualization in the order of a billion rows per second, for a high-end desktop computer. We use streaming algorithms, memory mapped files, and a zero memory copy policy to allow exploration of datasets larger than memory, for example out-of-core algorithms. Vaex allows arbitrary (mathematical) transformations using normal Python expressions and (a subset of) numpy functions which are “lazily” evaluated and computed when needed in small chunks, which avoids wasting of memory. Boolean expressions (which are also lazily evaluated) can be used to explore subsets of the data, which we call selections. Vaex uses a similar DataFrame API as Pandas, a very popular library, which helps migration from Pandas. Visualization is one of the key points of vaex, and is done using binned statistics in 1d (e.g. histogram), in 2d (e.g. 2d histograms with colourmapping) and 3d (using volume rendering). Vaex is split in in several packages: vaex-core for the computational part, vaex-viz for visualization mostly based on matplotlib, vaex-jupyter for visualization in the Jupyter notebook/lab based in IPyWidgets, vaex-server for the (optional) client-server communication, vaex-ui for the Qt based interface, vaex-hdf5 for HDF5 based memory mapped storage, vaex-astro for astronomy related selections, transformations, and memory mapped (column based) FITS storage.
Key words: methods: numerical / methods: statistical
The interactive image associated to Fig. 4 is available at https://www.aanda.org
Vaex is open source and available under MIT license on github, documentation and other information can be found on the main website: https://vaex.io or https://github.com/maartenbreddels/vaex
© ESO 2018
1. Introduction
Visualization plays an important role in astronomy, and is often used to find and display trends in data in the form of two dimensional scatter plots. The Herzsprung-Russel diagram for example, is one of the most famous scatter plots, showing the relationship between the temperature and the luminosity of stars. Before the era of computers, these plots were drawn by hand, while now it is customary to use a software package or a library to produce them.
While two-dimensional scatter plots may reveal trends or structure in a dataset of relatively small size, they become illegible when the number of samples exceeds ∼ 106: the symbols overlap and ultimately fill up the plot in a uniform colour, obscuring any information such a figure may contain. When a dataset contains more than 106 samples, it is more meaningful to visualize the local density in a two dimensional plot. The density can be determined using a kernel density estimator (KDE) or by a binning technique, equivalent to constructing a histogram in one dimension. The value of the local density can then be translated to a colour using a colourmap, which makes for an informative visualization.
To illustrate this concept, in Fig. 1 we show the positions in equatorial coordinates of the stars in the Gaia DR1 catalogue (Gaia Collaboration 2016), which contains over 1 billion entries in total. On the top left panel we show a scatter plot containing only 104 randomly chosen stars. This plot shows some structure, the Galactic disk is clearly seen, and one can also see the Large Magellanic Cloud as an over-density of points at (RA, Dec) ≈ (80°, −70°). These structures are largely smeared out and nearly unnoticeable on the right panel where we show a scatter plot with 1 000 000 stars. On the other hand, we obtain significantly more information if we visualize the data with a density plot. The bottom panel in Fig. 1 shows a density plot of the entire Gaia DR1 catalogue, where one can see in great detail the structure of the disk, the Magellanic Clouds, patterns related to how the satellite scans the sky, and even some dwarf galaxies and globular clusters. All these details are lost when we represent the data with a scatter plot. However, instead of providing a pure visualization library, the visualization is part of a larger library that provides efficient transformation, filtering and storing of the data, as well as efficient algorithms to calculate statistics that form the basis of the visualization.
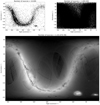 |
Fig. 1. Comparison between scatter and density plots when trying to visualize the sky as observed by Gaia DR1. Top left: a scatter plot showing 10 000 sources in Equatorial coordinates, which does reveal some structure in the disk. Top right: idem, with 1 000 000 sources, hiding almost any structure present in the data. Bottom: a density plot with 1 142 679 769 sources, where the logarithm of the density is colour-mapped (black is low density and white is high density). This reveals much more features in the data, such as structure in the disk and artefacts related to the scanning nature of the Gaia satellite. |
In this paper we present a new Python library called vaex, which is able to handle extremely large tabular datasets such as astronomical catalogues, N-body simulations or any other regular datasets which can be structured in rows and columns. Fast computations of statistics on regular N-dimensional grids allows analysis and visualizations in the order of a billion rows per second. We have used streaming algorithms, memory mapped files and a zero memory copy policy to allow exploration of datasets larger than the random-access memory (RAM) of a computer would normally allow, for example out-of-core algorithms. Vaex allows arbitrary mathematical transformations using normal Python expressions and numpy functions which are lazily evaluated, meaning that they are only computed when needed, and this is done in small chunks which optimizes the RAM usage. Boolean expressions, which are also “lazily” evaluated, can be used to explore subsets of the data, which we call selections. Vaex uses a similar DataFrame API as Pandas (McKinney 2010), a very popular Python library, which lessens the learning curve and makes its usage more intuitive. Visualization is one of the focus points of vaex, and is done using binned statistics in one dimension (histogram), in two dimensions (2d histograms with colour-mapping) and in three dimensions (using volume rendering). Vaex is split into several packages: vaex-core for the computational part, vaex-viz for visualization mostly based on matplotlib, vaex-server for the optional client-server communication, vaex-ui for the Qt based interface, vaex-jupyter for interactive visualization in the Jupyter notebook/lab based on IPyWidgets, vaex-hdf5 for HDF5 based memory mapped storage and vaex-astro for astronomy related selections, transformations and col-FITS storage. Vaex is supported on Linux, OSX and Windows.
Other similar libraries or programs exist, but do not match the performance or capabilities of vaex. TOPCAT (Taylor 2005), a common tool in astronomy, has support for density maps, but in general is focussed on working on a per row basis, and does not handle 109 objects efficiently. Glue (Beaumont et al. 2015; Robitaille et al. 2017) is a similar tool, but also allows mixing of different type of data, tabular and image data. The Pandas library can be used for similar purposes, but its focus is on in memory data structures. The datashader1 library can handle large volumes of data, focuses mainly on visualization in two dimensions, and lacks tools for exploration of the data. Dask2 and especially its DataFrame library is a good alternative for the computational part of vaex but it is accompanied with a steeper learning curve, and inherits the memory and performance issues of Pandas. The Gaia archive hosts a visualization page specialized for Gaia (Moitinho et al. 2017), which allows many users to efficiently explore pre-defined visualization.
This paper is structured as follows. In Sect. 2 we begin by laying out the main ideas that form vaex, which we support with the relevant calculations. In Sect. 3, we first present the basis of the library (vaex-core), and discuss all other packages that are subsequently built on top of it, such as vaex-astro for the astronomy related package, and vaex-ui for the Qt based user interface. We summarize our work in Sect. 4.
We note that this paper does not document all the features and options of the the vaex library. It lays out the principle ideas and motivation for creating the software, and presents the main capabilities of vaex3. Vaex is open source and available under the MIT licence on github4 or Zenodo5.
2. Motivation for development of Vaex
In this section, we lay out the main ideas and the motivation for developing vaex. We start by discussing the possibilities and limitations when dealing with large tabular datasets. With a simple calculation we show that it should be theoretically possible to process one billion samples per second, and demonstrate this with an implementation.
2.1. Constraints and possibilities
In the Introduction we showed a clear example of how a scatter plot displaying ∼109 samples is usually not meaningful due to over-plotting of the symbols. In addition, when one wants to process 109 samples in one second on a machine with an Intel(R) Core(TM) i7-4770S CPU 3.1 GHz CPU with four cores6, only 12.4 CPU cycles are available per sample. That does not leave room for plotting even one glyph per object, as only a few CPU instructions are available. Furthermore, considering numerical data for two columns of the double precision floating point type, the memory usage is 16 GB (109 × 2 × 8 bytes = 16 × 109 bytes ≈ 15 GiB), which is a significant fraction of the maximum bandwidth of 25.6 GB s−1 for this CPU.
Therefore, for the vaex library we only consider streaming or out-of-core algorithms which need one or a few passes over the data, and require few instructions per sample. The computation of statistics such as the mean or higher moments of the data are examples of such algorithms. The computation of a histogram on a regular grid can also be done with only few instructions, enabling us to efficiently visualize large amounts of data in one dimension by means of a histogram, in two dimensions via a density plot, and in three dimensions by the means of volume or isosurface rendering.
Preprocessing the data can lead to additional increase in performance. However, we assume that users may wants to perform various transformations on the data while they are exploring it, such as taking the log of a quantity or the difference between two quantities. Although some likely transformations can be provided, the number of combinations in combination with not knowing all possible transformations beforehand, we do not consider preprocessing feasible.
2.2. Real performance
We implemented a simple binning algorithm in C with a Python binding, finding that we can create a 256 × 256 two dimensional histogram from a dataset with 0.6 billion samples in 0.55 s, corresponding to a processing rate of 1.1 billion objects s−1, which we consider acceptable for interactive visualization. This code uses multi-threading7 to achieve this speed, while using ∼75−85% (15–17 GB s−1) of the maximum memory bandwidth8.
We note that in these calculations, the data was already cached in the operating system’s cache. If we would purge the cache, or choose two columns that were not already in the cache it would take for this example 90 s for a slow spinning hard drive at 100 MB s−1, to less than 3 s for an SSD that can reach 3.2 GB s−1 sequential read speeds.
2.3. N-dimensional statistics
Apart from simply counting the number of samples in each bin, one can generalize this idea to calculate other statistics per bin using extra columns. Instead of simply summing up the number of samples that fall into each bin, one can use the same algorithm to perform other computations on a particular column, effectively calculating many statistics on a regular grid in N dimensions, where 0 dimensions implies a scalar. For example, let us consider a dataset that features four columns x, y, υx, υy, where the first two represent the position and the last two the corresponding velocity components of a particle or a star. One can construct a two dimensional grid spanned by x and y displaying the mean υx by first summing up the υx values and then dividing by the total number of samples that fall into each bin. The same exercise can be repeated to calculate the mean velocity in y direction. Higher order moment can also be calculated, allowing one to compute and visualize vector and tensor quantities in two and three dimensions. The types of statistics available in vaex are listed in Sect. 3.1.6 and Table 2.
2.4. Implementation
These ideas and algorithms, which are efficiently implemented, form the basis of the vaex library. Vaex exposes them in a simple way, allowing users to perform computations and scientific visualizations of their data with minimal amount of code. The graphical user interface (GUI) application, uses the library to directly visualize datasets to users, and allows for interactive exploration. By this we mean the user is not only able to navigate (zoom and pan), but also to make interactive selections (visual queries), which can be viewed in other windows that display a plot created by a different combination of columns from those on which the selection was made on (linked views). Vaex also provides ranking of subspaces9, by calculating their mutual information or correlation coefficient in order to find out which subspaces contain more information.
2.5. Using a subset of the data
In some cases, it may be useful to do computations on a smaller random subset of of data. This is beneficial for devices that do not have enough storage to keep the whole dataset such as laptops, and will also require less computing power. This is also useful for servers, as we will see in Sect. 3.7 in order to handle many requests per second. Instead of drawing a random subset of rows from the full dataset, we store the dataset with the rows in a random order, and than “draw” a random subset of rows (which will be the same every time), by only processing the first N rows. To support this, the library includes the option to convert and export a dataset with the rows in a random order.
3. Vaex
The ideas of the previous section form the basis of the vaex library. Vaex is a Python (meta) package, consisting of a several Python packages, where vaex-core also contains a so called extension module, written in C, which contains the fast performing algorithms, such as those for binning the data. The vaex package can be installed using pip, or (ana)conda10. Its source code and issue tracker are on-line11.
Installing vaex (the meta package) will install all packages in the vaex family. However, if only a few functionalities are needed, only the relevant packages can be installed. For instance in many cases only vaex-core, vaex-hdf5 and vaex-viz are needed. One can thus avoid installing vaex-ui since it has (Py)Qt as a dependency, which can be more difficult to install on some platforms. Table 1 lists all packages with a short description.
Packages in the vaex family.
3.1. vaex-core
The foundation of all vaex packages is vaex-core. This contains the most important part, the Dataset class, which wraps a series of columns (numpy arrays) in an API similar to Pandas’ DataFrames, and gives access to all the operations that can be performed on them, such as calculating statistics on N-dimensional grids or the joining of two tables. On top of that, the Dataset class does bookkeeping to track virtual columns, selections and filtering. Note that in vaex almost no operation makes memory copies of the data (unless explicitly requested, or required), since we expect the full dataset to be larger than the RAM of typical computer.
3.1.1. (Lazy) expressions
In practice, one rarely works only with the columns as they are stored in the table. Within the vaex framework, every statistic is based on a mathematical expression, making it possible to not just plot the logarithm of a quantity for example, but to plot and compute statistics using an arbitrary, often user defined expression. For instance, there is no difference in usage when calculating statistics on existing columns, for example the mean of x, or any mathematical operation using existing columns, for example x + y, where x and y are two columns of a vaex dataset. The last expression will be calculated on-the-fly using small chunks of the data in order to minimize memory impact, and optimally make use of the CPU cache. Being able to calculate statistics on an N-dimensional grid for arbitrary expressions is crucial for exploring large datasets, such as the modern astronomical catalogues or outputs of large-scale numerical simulations. For instance taking the logarithm of a column is quite common, as well as calculating vector lengths (e.g. 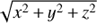 ). No pre-computations are needed, giving users the complete freedom of what to plot or compute.
). No pre-computations are needed, giving users the complete freedom of what to plot or compute.
In the the commonly used Pandas library, a statement like a = df.b + np.sin(df.c), where df is a Pandas DataFrame containing the columns b and c, would be directly computed and will result in additional memory usage equal to that of the columns b or c. In constrast, the statement a = ds.b + np.sin(ds.c), where ds is a vaex Dataset, results in an expression which only stores the information of how the computation should be done. The expression will only be calculated when the result of a is needed, which if often referred to as a lazy evaluation. For convenience, a vaex Dataset can also hold what we refer to as virtual columns, which is a column that does not refer to a numpy array, but is an expression. This means that many columns can be added to a dataset, without causing additional memory usage, and in many cases causing hardly any performance penalty (when we are not CPU-bound). For instance, a vector length can be added using ds[“r”] = np.sqrt(ds.x**2 + ds.y**2 + ds.z**2), which can then be used in subsequent expressions. This minimizes the amount of code that needs to be written and thus leads to fewer mistakes.
3.1.2. Just-in-time compilation
Once the result of an expressions is needed, it is evaluated (in blocks) using the numpy library. For complex expressions this can result in the creation of many temporary arrays12, which may decrease performance. In these cases, the computation speed of such complex expressions can be improved using just-in-time (JIT) compilation by utilizing the Pythran (Guelton et al. 2015) or Numba (Lam et al. 2015) libraries which optimize the code at runtime. Note that the JIT compilation will not be done automatically, but needs to be manually applied on an expression, e.g. ds[“a”] = (ds.x + ds.y + ds.z * ds.w).jit().
3.1.3. Selections and filtering
In many cases one wants to visualize or perform computations on a specific subset of the data. This is implemented by carrying out so called selections on the dataset, which are one or more boolean expressions combined with boolean operators. Such selections can be defined in two ways, via boolean expressions, or via a geometrical (lasso) selection. The boolean expressions have the same freedom as expressions applied on the dataset when computing statistics, and can be a combination of any valid logical expressions supported by Python using one or more (virtual) columns. Examples of such booleans expressions are np.sqrt(ds.x**2 + ds.y**2 + ds.z**2) < 10 or (ds.FeH > −2) & (ds.FeH < −1), where the ampersand means logical “and”. Although the geometrical lasso selection can be implemented via boolean expressions, it is implemented separately for performance reasons. The lasso selection can be used a in graphical user interfaces to select regions with a mouse or other pointing device, or to efficiently select complex regions in two dimensions such as geographical regions. A dataset can have multiple selections, and statistics or visualizations can be computed for one or more selections at the same time (e.g. ds.mean(ds.x, selection = [ds.x < 0, ds.y < 0])) in a single pass over the data. When selections are created using “select” method ds.select(selection = ds.x < 0, name = “negative”), they can be named by passing a string to the name argument, and the result of the selection, which is a boolean array, will be cached in memory leading to a performance increase. If no name is given, it will assume the name “default”. Thus all selection arguments in vaex can take a boolean expression as argument, a name (referring to a selection made previously with ds.select), or a boolean, where False refers to no selection and True to the default selection. This is useful for selections that are computationally expensive or selections that are frequently used. In the current implementation, a named selection will consume one byte per row, leading to a memory usage of 1 GB for a dataset containing 109 rows. Note that no copies of the data are being made, only a boolean mask for the selection is created. Note that also selections are lazy, and will be computed in chunks only when needed, similar to expressions.
Often, a part of the data will not be used at all as part of preprocessing or cleaning up. In this case we want a particular selection to be always applied, without making a copy of the data. We refer to this as filtering, and is similarly done as in Pandas, e.g. ds_clean = ds[ds.x_error < 10]). The filtering feature is implemented in exactly the same way as the selections, except that a filter will always be applied, whereas a selection has to be passed to a calculation explicitly each time.
A history of the expressions that define the selections is also kept, which leads to less memory usage and enables users to undo and redo selections. In addition, the expressions which define a selection can be stored on disk, making the steps that led to a particular selection traceable and reproducible.
3.1.4. Missing values
It often happens that some samples in a column of a dataset lack an entry. Such missing values are supported using numpy’s masked array type, where a boolean is kept for every row of a column, specifying whether a value is missing or not. For floating point numbers a NaN (Not a Number) values are also interpreted as missing values. Masked arrays are important for integer data, which does not have a NaN equivalent. Both masked values and NaN’s are treated as missing values for all the statistical calculations.
3.1.5. Units
Optionally, a unit can be assigned to a column. Expressions based on columns which all have units assigned to them will also result is an unit for that expression. For visualization, the units can be used in the labelling of axis, as is done in vaex-viz (see the x-axis labelling of the top panel of Fig. 2 for an example). The use of units fully relies on the core Astropy Python package (Astropy Collaboration 2013).
 |
Fig. 2. Example of one dimensional visualization for the vaex-viz package. Top: histogram of Lz, the angular momentum in the z direction. Because the units are specified in the data file for this column, it will be included in the labelling of the x axis by default. Bottom: similar as above, but showing the mean of the energy E in each bin. |
3.1.6. Statistics on N dimensional grids
One of the main features of vaex is the calculations of statistics on regular N-dimensional grids. The statistics, listed in Table 2 can be computed for zero13 or higher dimensional grids, where for each dimension an expression, a bin-count and a range (a minimum and a maximum) must be specified. All these calculations make use of the fast N-dimensional binning algorithm which is explained in more detail in Appendix A. Each method can take one or more selections as arguments, which will then stream over the data once, or as few times as possible, giving optimal performance especially when the size of the data exceeds that of the RAM.
Functions which can be computed on N dimensional grids and exploit the fast binning algorithm, which are readily available in vaex.
3.2. vaex-hdf5
In order to achieve the performance estimated in Sect. 2, we need to put some constraints on how the data is stored and accessed. If we assume the data is in the operating system cache, and we want to access the data using a normal read call, this will result in a memory copy from the OS’ cache to the array. Alternatively, if the data is stored in a file format that can be memory mapped, it will enable us to directly use the physical memory of the operating system cache, eliminating unnecessary overheads. Aside from the memory mapping requirements, we also impose additional constrains on the file format in which the data is to be stored. First, we require the data to be stored in the native floating point format of the CPU (IEEE 754), and preferably in the native byte order (little endian for the x86 CPU family). Our second requirement is that the data needs to be stored in a column based format, meaning that the datum of the next row for a given column is in the next memory location. In cases where we only use a few columns, such as for visualization, reading from column based storage is optimal since the reading from disk is sequential, giving maximum read performance.
The well known and flexible file format HDF5 has the capabilities to do both column based storage and to store the data in little and in big endian formats. The downside of the HDF5 format is that it can store almost anything, and there are no standards for storing meta information or where in the file a table should be stored. To reconcile this issue, we adopted the VOTable metadata naming and convention (Ochsenbein et al. 2013), including support for Unified Content Descriptors Derriere et al. (2004, UCD), units and descriptions for every column, and a description for the tables. Having UCDs and units as part of the column description allows the software to recognize the meaning of the columns, and suggest appropriate transformations for example. The layout of the file is explained in more detail online14.
Although vaex can read other formats, such as FITS, ascii or VOTable, these require parsing the file or keeping a copy in memory, which is not ideal for datasets larger than ≳100 MB. For superior performance, users can convert these formats to HDF5 using vaex. An intermediate solution is the column based FITS format, which we will discuss in Sect. 3.6.
3.3. vaex-viz
A core feature of vaex is the visualization based on statistics calculated on N-dimensional grids. The vaex-viz package provides visualization utilising the matplotlib library (Hunter 2007).
To showcase the main feature of vaex, we use a random 10% subset of the dataset generated by Helmi & de Zeeuw (2000) which will be downloaded on the fly when ds = vaex.example() is executed. This dataset is a simulation of the disruption of 33 satellite galaxies in a Galactic potential. The satellites are almost fully phase-mixed, making them hard to disentangle in configuration space15, but they are still separable in the space spanned by the integrals of motion: E (Energy), L (total angular momentum)16, and the angular momentum around the z axis Lz. Even though this dataset contains only 330 000 rows, it serves well to demonstrate what can be done with vaex while being reasonably small in size.
Larger datasets, such as the full Helmi & de Zeeuw (2000) dataset, the Gaia DR1 and DR2 catalogues (Gaia Collaboration 2016), or over one billion taxi trips in New York can be found online17.
For one dimensional visualizations, all statistics listed in Table 2 can be plotted as a function of a single parameter or an expression, where the count method will be used to create a histogram. An example is shown in Fig. 2, where on the top panel we show a regular one dimensional histogram for Lz, while in the bottom panel we visualize the mean energy E in each bin in log radius. Note that the x-axis in the top panel include a unit by default because these are include in the data file.
For two dimensional visualizations, we can display a two dimensional histogram as an image. An example of this is shown in the top left panel of Fig. 3, which shows a plot of y versus x (the positions of the simulated particles) where the logarithm of the bin counts is colour-mapped. Again, note the units that are included by default on the axes. Similarly as for the one dimensional case, we can also display other statistics in bins as shows in the top right panel of Fig. 3. Here instead of showing the counts in bins, we display the standard deviation of the z component of the velocity vector (υz). This already shows some structure: a stream that is not fully phase mixed can be readily seen.
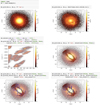 |
Fig. 3. Example of two dimensional visualization for the vaex-viz package. Top left: two dimensional histogram, colour coded by a colour map visualizing the counts in each bin. Top right: similar to the left, but showing the standard deviation in each bin. Middle left: a zoom-in of the E versus Lz space, where a rectangular region is selected using vaex’ selection mechanism. The underlying visualization of all the data is slightly faded out to highlight the selection. Middle right: similar to the top left, but displaying the both the full dataset and the selection, where the selected rectangular region corresponds to a satellite that is not fully phase mixed. Bottom left: similar as the middle right plot but overlaying the mean velocity of the selection using vectors. Bottom right: similar as the bottom left, except now overlaying the velocity dispersion tensor represented by ellipses. |
In the middle left panel of Fig. 3, we create a selection (a rectangle in E and Lz space), and visualize the full dataset and the selection on top of each other. The default behaviour is to fade out the full dataset a bit so that the last selection stands out. Since selections are stored in the Dataset object, subsequent calculations and plots can refer to the same selection. We do this in the right panel of the middle row of the same figure, where we show in a different space (y vs. x) what the full dataset and the selection look like. Here we can clearly see that the clump selected in E and Lz space corresponds to a not fully phase-mixed stream, the same structure we noticed in the top right panel in this figure.
On the bottom left panel of Fig. 3, we use the same selection but a different visualization. We first repeat the same visualization as in the middle right panel, but overlay vectors, or a quiver plot. The plot2d_vector method calculates a mean vector quantity on a (coarser) grid, and displays them using vectors. This can give better insight into vectorial quantities than for instance two separate density plots. The grid on which the vectorial quantity is computed is much coarser to not clutter the visualization. Optionally, one can also colour-map the vectors in order to display a third component, for instance the mean velocity in the z direction. Using a combination of the density map and the vector fields, one can plot up to five different dimensions on the same figure.
Similarly to a vectorial quantity, we can visualize a symmetric two dimensional tensorial quantity by plotting ellipses. We demonstrate this on the bottom right panel in Fig. 3 using plot2d_tensor, where we visualize the velocity dispersion tensor of the x and y components for the same selection. In this visualization a diagonal orientation indicates a correlation between the two velocity dispersion components and the size of the ellipse corresponds to the magnitudes of the velocity dispersions.
3.4. vaex-jupyter
The vaex library, especially the visualization tools described in Sect. 3.3 can also be used in combination with the Jupyter (formerly IPython) notebook (Pérez & Granger 2007)18 or Jupyter lab, which is a notebook environment in the web browser. A web browser offers more options for interactive visualization and thus exploration of data compared to the static images which are the default in matplotlib inside the Jupyter notebook. The Jupyter environment allows vaex to work in combination with a variety of interactive visualization libraries, mostly built on ipywidgets19. For two-dimensional visualization bqplot20 allows for interactive zooming, panning and on-plot selections as shown in the top panel of Fig. 4. ipympl21 is an interactive back-end for matplotlib, but unlike bqplot, the visualization is rendered in the kernel as opposed to in the browser, giving a small delay when rendering a plot. The ipyleaflet22 library can be used to overlay and interact with a geographical map.
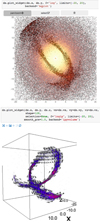 |
Fig. 4. Top: screenshot of a Jupyter notebook using the widget back-end to visualize the dataset interactively using panning, zooming and selection with the mouse. The back-end used here is bqplot. Bottom: a 3d visualization using the ipyvolume backend, showing the stream discussed before, where the vector visualize the mean velocity. An interactive version of this visualization is available online. |
To display the data in three dimensions we use ipyvolume23, which offers volume and isosurface rendering, and quiver plots using WebGL. A big advantage of using WebGL in the Jupyter notebook is that it allows one to connect to a remote server while running the visualization on the local computer, a feature that is difficult to set up using OpenGL24. The bottom panel of Fig. 4 shows an example of three dimensional volume rendering including a quiver plot, created using the synergy between vaex and ipyvolume in the Jupyter environment. We note that in the three dimensional version, especially the interactive version on-line, gives a much clearer view on the orientation and direction of rotation of the stream compared to the two dimensional version shown in Fig. 3. These backends for interactive two and three dimensional visualization are all implemented in the vaex-jupyter package.
3.5. vaex-ui
The vaex-ui package provides a graphical user interface that can be used as a standalone GUI application or used in the Jupyter environment. We focus now on the standalone GUI application. Upon starting, the application shows the open tables on the left side, as shown in the top left panel on Fig. 5. On the right side it shows metadata information of the selected table, and it contains buttons for opening windows that can visualize the dataset, give statistics on the data, show the ranking of the subspaces, or display the whole table. The next tabs shows the available columns, allows editing of units and UCDs, and addition and editing of virtual columns. The third tab shows the variables that can be used in expressions.
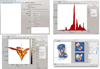 |
Fig. 5. Top left: a screenshot of main application window. On the left it shows the open tables, and on the right the meta-data and operations that can be performed on the table. Top right: one dimensional histogram showing the Lz distribution of our example dataset. Bottom left: a two dimensional plotting window, showing E versus Lz. Bottom right: a volume rendering of a stream in our example dataset in Cartesian coordinates. The mean velocity field is also displayed with the help of vectors. |
As in Sect. 3.3, one can do one dimensional visualization using histograms or statistics in regular bins. The top right panel in Fig. 5 shows an example of this, where we plot the histogram of Lz for the example dataset as presented in Sect. 3.3. The plot shown on this figure is interactive, allowing zooming and panning with the mouse and keyboard, as well as interactive selections. In the “x-axis” text box of this window, one can enter any valid mathematical Python expression, such as log(sqrt(x**2 + y**2 + z**2)) for example, where x, y and z are columns in the dataset we use. In addition to the columns present in the dataset, one can also use any pre-defined virtual columns or variables to compute user defined expressions. Apart from the standard histograms that are based on the count statistic in Table 2, users can visualize other statistics, such as the mean or standard deviation of an expression per bin.
For two dimensional visualizations, the GUI application displays a two dimensional histogram using an image, also similar to Sect. 3.3. An example of this is shown in the bottom left panel of Fig. 5, which is a plot of the E versus the Lz, where the logarithm of the bin counts is colour-mapped. On this panel, one can see the individual satellites, each having its own distinct energy and angular momentum. As in the case for the one dimensional histogram, the entries for the x- and y-axis fields can be any valid mathematical Python expressions. Also similar to the one dimensional visualization, the statistics listed in Table 2 can be visualized in the bins, now in two dimensions. Vectorial and symmetric tensor quantities can be displayed as well, in a similar manner to what is described in Sect. 3.3.
The GUI application also supports volume rendering using the OpenGL shading language. We supports multi-volume rendering, meaning we can display both the full dataset and a GUI application also selection. In addition, one can over-plot vector fields in three dimensions. The users have access to custom settings for the lighting and for the transfer function. Navigation and selections are done in two dimensional projection plots displayed alongside the panel that shows the three dimensional rendering. An example of this visualization is shown in the bottom right panel of Fig. 5.
3.5.1. Linked views
The GUI application also supports linked views (Goodman 2012). This feature makes all the active plots of the application linked to a single selection, allowing for complex and interactive exploration of a dataset. To demonstrate this concept, the left panel of Fig. 6, shows a zoomed-in view of the bottom left panel of Fig. 5, where we have selected a particular cluster of stars in the subspaces spanned by E and Lz. This is similar to what we have done in the middle left panel of Fig. 3, except that we can now do the selection interactively with the mouse. On the right panel of Fig. 5 we see what the selection looks like in configuration space, and we can readily see that the selected clump in E and Lz space is in fact a stream. This is confirmed by the velocities of its constituents stars, which are displayed with the help of a vector field overlaid on the same panel.
 |
Fig. 6. Left: a two dimensional plot window, showing E vs. Lz, similar to the middle left panel of Fig. 3. Right: a two dimensional plot windows, showing x vs. y, similar to the middle right panel of Fig. 5, but sharing the same selection as that shown in the left panel, demonstrating the linked views feature. |
3.5.2. Common features
The windows that display the plots contain a number of options to aid users in the exploration and visualization of their data. These include the setting of an equal axis ratio, keeping a navigation history, undoing and redoing selections, and numerous options to adjust the visualization settings. Users also have the option to display multiple dataset on top of each other on the same figure using the concept of layers, where where different blending operations can be set by the user. There are also options for exporting figures as raster (e.g. jpeg) or vector (e.g. pdf) graphics, as supported by the matplotlib library. It is also possible to export the binned data and with it a script that reproduces the figure as seen on the screen. This enables users to further customize their plots and make them publication ready.
3.5.3. Subspace exploration/ranking
Many tables nowadays contain a large number of columns. Thus, inspecting how different quantities present in a dataset depend on each other, and finding which subspaces, or combinations of those quantities in two or more dimensions contain the most information via manual inspection by the user can be quite tedious, and sometimes not feasible at all. To aid users in finding subspaces that are rich in information, the GUI application offers the option to rank subspaces according to two metrics: the Pearson’s correlation coefficient and the mutual information. The calculation and interpretation of the Pearson’s correlation coefficient is well documented in classical statistics, and we will not discuss it further.
The mutual information is a measure of the mutual dependence between two or more random variables. It measures the amount of information one can obtain about one random variable through the measurement or knowledge of another random variable. In vaex, the mutual information is obtained using the Kullback–Leibler Divergence (Kullback & Leibler 1951, KLD), and for two random variables it is calculated via the expression:
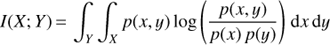 (1)
(1)
where p(x, y) is the joint probability distribution of the random variables x and x, while p(x) and p(y) are their marginalized probability distributions. This quantity can be calculated for all, or a user defined subset of subspaces having two or more dimensions. The mutual information effectively measures the amount of information contained within a space spanned by two or more quantities. On the left panel of Fig. 7 we show the window listing all two dimensional subspaces in our example dataset, sorted by mutual information. In the right panel of this figure we show two subspaces that have the highest and two subspaces that have the lowest rank according to their mutual information, in the top and bottom rows respectively. One can readily see that the spaces spanned by the integrals of motion (E, L and Lz) are found to contains the most information, as expected.
 |
Fig. 7. Left: ranking of subspaces my mutual information. Right, top row: two subspaces having the highest; bottom row: two subspaces with the lowest mutual information rank. |
3.6. vaex-astro
This subpackage contains functionality mostly useful for astronomical purposes.
3.6.1. FITS
Although FITS is not as flexible as HDF5, and is not designed to store data in column based format, it is possible to do so by storing one row where each column contains a large vector. TOPCAT is using this strategy, and calls it the col-FITS format. However, the BINTABLE extension for FITS mandates that the byte order to be big endian. This mean that the bytes need to be swapped before use, which in vaex gives a performance penalty of ∼30%. For compatibility with existing software such as TOPCAT, we support the col-FITS format both for reading and writing.
3.6.2. Common transformation
The conversion between different coordinate systems is common in astronomy. The vaex-astro package contains many of the most common coordinate transformations, such as the conversion between positions and proper motions between different spherical coordinate systems (Equatorial, Galactic, Ecliptic), as well as the conversion of positions and velocities between spherical, Cartesian and cylindrical coordinate systems. The transformations also include the full error propagation of the quantities in question. If the dataset contains metadata describing the units of the quantities, these will be automatically transformed as needed, minimizing possible sources of error.
3.6.3. SAMP
Vaex offers interoperability with other applications via the Simple Application Messaging Protocol (SAMP Boch et al. 2009). It can communicate with a SAMP hub, for instance by running TOPCAT’s build in SAMP hub, which can then broadcast a selection or objects that are “picked” to other applications. Vaex understand the “table.load.votable” message, meaning other applications can send tabular data in the VOTable format to it. Although this transfer mechanism is slow, it means that any data that can be read into TOPCAT can be passed on to vaex. For example, one can download a VOTable from VizieR using the TAP protocol (Dowler et al. 2010), which is fully implemented in TOPCAT, and than use SAMP to transfer it to vaex. The same is possible with any other application or web service that supports SAMP.
3.7. vaex-server
A dataset that consists of two columns with 1 billion rows filled with double precision floating point values amounts to 16 GB of data. On the other hand, if this dataset is binned on a 256 × 256 grid, which also uses double precision floating points for its bin values, has the size of only 0.5 MiB. Using a suitable compression this can be reduced further by a factor of approximately 10. This makes it possible and practical to do calculations on a server and transfer only the final results to a client.
A working client/server model is implemented in the vaex-server package, making it possible to work with both local and remote datasets at the same time. Also the GUI application provided by vaex-ui allows a users to connect to a remote server. The server itself is completely stateless, meaning it does not keep into memory the state of a remote client. This means that when a user requests a statistic for a subsets of the data defined by a selection, the client needs to send that selection to the server, and the server will compute the selection at each request. The server can cache selections, speeding up subsequent calculations of statistics for which the same subset of the data is required. The benefit of having it stateless is that the server can be less complex, and can be scaled vertically, meaning that more servers can be added with a load balancer in front to scale up the workload.
Furthermore, clients have the option to let the server determine how much of the data it will use, assuming the data is shuffled, to give a approximate answer. The server will then estimate how much of the data should be processed to return a result in a predefined amount of time, which is one second by default. Clients that want to use a specific amount of data, up to 100%, may need to wait longer. Using this mechanism, the vaex-server can handle up to 100 requests per second on a single computer.
3.8. vaex-distributed
Many of the statistics that vaex can compute, such as the mean or variance can be written as a linear combination of smaller chunks of the data used. Thus it is possible to distribute the computations to different computers, each performing a part of the work on a smaller data chunk, and combine the results at the end. The vaex-distributed package makes this possible. With its use, we manage to perform computations 10 billion (1010) rows per second on a cluster of 16 low-end computers. In this case we managed to overcome the memory bandwidth bottleneck, something that could not have been solved by having a single computer with the same number of cores as the cluster. This demonstrates that vaex can be scaled up to even larger datasets of the order of ≈10−100 billion rows with the help of a computer cluster, even if such a cluster is not composed of modern computers. Note that not all functionality is supported, as this is only a proof of concept.
3.9. vaex-ml
Build on top of vaex, is another proof of concept package called vaex- ml, which combines machine learning with vaex’ efficient data handling. Comparing vaex-ml a k-means clustering algorithm to that of scikit- learn (Pedregosa et al. 2011), we are about 5× faster and have low memory impact since vaex does not need to load the dataset in memory nor copies it. In both cases all cores are used. For PCA we are almost 7× faster, but vaex by default uses multi-threading for the calculation of the covariance matrix. Note that the sklearn implementations of both PCA and k-means are limited to 10 million rows in order to avoid swapping memory to disk, while vaex happily works through the 100 million rows. We furthermore integrated vaex with xgboost (Chen & Guestrin 2016) to make boosted tree models easily available. Note that the vaex-ml source code is available and is free for personal and academic usage.
3.10. Any large tabular dataset
We would like to emphasize that, even though the main motivation for creating vaex was to visualize and explore the large Gaia catalogue, vaex is an ideal tool to use when working with any other large tabular dataset. To illustrate this point, in Fig. 8 we visualize three large datasets. The leftmost panel is a density plot of approximately one billion drop-off locations made by the Yellow cab taxies in New York City for the period between 2009 and 2015. The middle panel on the same figure shows the positions for 0.6 billion particles from the pure dark matter Aquarius simulation (Aq-A, level 2 Springel et al. 2008). The right panel in Fig. 8 displays the Open street map GPS data over Europe, made by approximately two billion GPS coordinates. These plots demonstrate that vaex is a suitable tool for exploration and visualization of any large tabular datasets regardless whether they are related to astronomy or not.
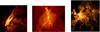 |
Fig. 8. Examples of density maps created from big datasets. Left: drop-off locations of the Yellow cab taxies in New York City between 2009 and 2015. In total ∼1 billion GPS coordinates are plotted. Middle: density plot of particle position from the Aquarius pure dark matter simulations (Springel et al. 2008, Aquarius A level 2), ∼0.6 billion particles. Right: open street map GPS data, showing ∼2 billion coordinates. |
4. Conclusions
In the future datasets will grow ever larger, making the use of statistics computed on N-dimensional grids for visualization, exploration and analysis more practical and therefore more common. In this paper we introduced vaex, a tool that handles large datasets, and processes approximately one billion rows per second on a single computer. The vaex Python library has a similar API to that of Pandas, making for a shallow learning curve for new users, and a familiar, easy to understand style for more experiences users. Built on top of many of the vaex packages is vaex-ui, which provides a standalone GUI application allowing data visualization in one, two and three dimensions, interactive exploration of the data such as panning and zooming, and visual selections. By combining the vaex GUI application, which can be used for a quick look at the data, with the vaex library for more advanced data mining using custom computations, users can be quite flexible in the manner in which they explore and make sense of their large datasets.
In the era of big data, downloading a large dataset to a local machine may not always be the most efficient solution. Using vaex-server, a dataset can be worked on remotely, only sending the final products, such as the (binned) statistics to the user. Combining multiple server in vaex-distributed allows vaex to scale effortless to ∼10 billion rows per second on small cluster of a dozen computers.
We have demonstrated many of the features of vaex using the example dataset from Helmi & de Zeeuw (2000). Visualization of statistics in one, two and three dimensions, using the full and subsets (selections in vaex) of the data as well as the visualization of vectorial and (symmetric) tensorial quantities. All of these calculations and visualizations will scale to datasets with billions of rows making vaex the perfect tool for the Visualization And EXploration (vaex) of the Gaia catalogue (Gaia Collaboration 2016), and even more so for the upcoming data releases or future missions such as LSST. The first data release of Gaia is available in HDF5 format online25, and we plan to do so as well for the second data release.
Vaex is open source, and available under the MIT license. Contributions are welcome by means of pull requests or issue reports26.
Interactive Figure
The interactive image associated to Fig. 4 Access Supplementary Material
The full documentation can be found at: https://vaex.io.
DOI: 10.5281/zenodo.1209875
This was considered a high-end desktop CPU, launched in the 2nd quarter of 2013.
Releasing Python’s Global Interpreter Lock when entering the C part to actually make use of the multi-threading.
Although the theoretical bandwidth is 25 GB s−1, we measured it to be 20 GB s−1 using the bandwidth program from http://zsmith.co/bandwidth.html.
We refer to a combination of one or more columns (or expression using columns) a subspace.
A popular Python distribution: https://www.continuum.io/downloads
https://github.com/maartenbreddels/vaex, and the home-page is at https://vaex.io.
Although from Numpy 1.13 temporaries can be avoided (Temporary elision), this is not guaranteed, and will still impact performance when it will not fit into L2 cache.
A scalar, or single value.
The 3d positions.
Although L is not strictly an integral of motion in an axi-symmetric system, see Helmi & de Zeeuw (2000).
Jupyter is the new front end to the IPython kernel.
IPython widgets, https://ipywidgets.readthedocs.io
Often, using remote X rendering falls back to using software rendering.
on https://github.com/maartenbreddels/vaex. The main website for vaex is https://vaex.io
Acknowledgments
MB and JV thank Amina Helmi for making this work possible. MB thanks Yonathan Alexander for pushing me to create a more user friendly API. MB thanks Anthony Brown for his support in the early stages of vaex. MB and JV AH are grateful to NOVA for financial support. This work has made use of data from the European Space Agency (ESA) mission Gaia (http://www.cosmos.esa.int/gaia), processed by the Gaia Data Processing and Analysis Consortium (DPAC, http://www.cosmos.esa.int/web/gaia/dpac/consortium). Funding for the DPAC has been provided by national institutions, in particular the institutions participating in the Gaia Multilateral Agreement.
References
- Astropy Collaboration, Robitaille, T. P., Tollerud, E. J., et al. 2013, A&A, 558, A33 [NASA ADS] [CrossRef] [EDP Sciences] [Google Scholar]
- Beaumont, C., Goodman, A., & Greenfield, P. 2015, ASP Conf. Ser., 495, 101 [Google Scholar]
- Boch, T., Fitzpatrick, M., Taylor, M., et al. 2009, Simple Application Messaging Protocol SAMP v1.11, IVOA [Google Scholar]
- Chen, T., & Guestrin, C. 2016, in Proc. 22nd ACM SIGKDD Int. Conf. Knowledge Discovery and Data Mining, KDD ’16 (New York, USA: ACM), 785 [CrossRef] [Google Scholar]
- Derriere, S., Gray, N., Mann, R., et al. 2004, (Unified Content Descriptor UCD) – Moving to UCD1+, IVOA [Google Scholar]
- Dowler, P., Rixon, G., & Tody, D. 2010, Table Access Protocol v1.0, IVOA [Google Scholar]
- Gaia Collaboration (Brown, A. G. A., et al.) 2016, A&A, 595, A2 [NASA ADS] [CrossRef] [EDP Sciences] [Google Scholar]
- Goodman, A. A. 2012, Astron. Nachr., 333, 505 [NASA ADS] [CrossRef] [Google Scholar]
- Guelton, S., Brunet, P., Amini, M., et al. 2015, Comput. Sci. Discovery, 8, 014001 [NASA ADS] [CrossRef] [Google Scholar]
- Helmi, A., & de Zeeuw, P. T. 2000, MNRAS, 319, 657 [NASA ADS] [CrossRef] [Google Scholar]
- Hunter, J. D. 2007, Comput. Sci. Eng., 9, 90 [NASA ADS] [CrossRef] [Google Scholar]
- Kullback, S., & Leibler, R. A. 1951, Ann. Math. Statist., 22, 79 [CrossRef] [MathSciNet] [Google Scholar]
- Lam, S. K., Pitrou, A., & Seibert, S. 2015, in Proc. Second Work-shop on the LLVM Comp. Infr. in HPC, LLVM ’15 (New York, USA: ACM), 7:1 [Google Scholar]
- McKinney, W. 2010, in Proc. 9th Python in Science Conference, eds. S. van der Walt, & J. Millman, 51 [Google Scholar]
- Moitinho, A., Krone-Martins, A., Savietto, H., et al. 2017, A&A, 605, A52 [NASA ADS] [CrossRef] [EDP Sciences] [Google Scholar]
- Ochsenbein, F., Williams, R., Davenhall, C., et al. 2013, VOTable Format Definition, Version 1.3, IVOA [Google Scholar]
- Pedregosa, F., Varoquaux, G., Gramfort, A., et al. 2011, J. Mach. Learn. Res., 12, 2825 [Google Scholar]
- Pérez, F., & Granger, B. E. 2007, Comput. Sci. Eng, 9, 21 [Google Scholar]
- Robitaille, T., Beaumont, C., Qian, P., Borkin, M., & Goodman, A. 2017, glueviz v0.10: Multidimensional Data Exploration [Google Scholar]
- Springel, V., Wang, J., Vogelsberger, M., et al. 2008, MNRAS, 391, 1685 [NASA ADS] [CrossRef] [Google Scholar]
- Taylor, M. B. 2005, ASP Conf. Ser., 347, 29 [Google Scholar]
Appendix A: Binning algorihm
The binning algorithm in vaex is a generalization of a one dimensional binning algorithm to N dimensions. It also supports custom operations per bin, on top of simply counting the number of samples that a bin contains. The algorithm itself is written in C and C++. For presentation purposes, we rewrote it in pure Python and how it in Fig. A.1 below. We consider Python code to be equivalent to pseudo code and thus self explanatory. The example includes the calculation of the counts, the minimum and maximum statistics, as well as the standard deviation on a regular two dimensional grid.
 |
Fig. A.1. Python code equivalent of our generalized Nd binning code, but only for 2d for clarity. |
# This is equivalent code for the c code, but written in Python for readability # It is for 2d only, 0, 1, and >= 3 dimensional are a generalization of this # but more difficult to read import numpy def operation_count(input, aux): if aux is not None and numpy.isinf(aux): return input else: return input+1 def operation_minmax(input, aux): if numpy.isinf(aux): return input else: return min(input[0], aux), max(input[1], aux) def operation_moment_012(input, aux): if numpy.isinf(aux): return input else: return [input[0] + 1, input[1] + aux, input[2] + aux**2] def statistic2d(grid, x, y, aux, xmin, xmax, ymin, ymax, operation): grid_width, grid_height = grid.shape[:2] # get dimensions of the 2d grid for i in range(len(x)): # iterator over all rows # normalize the x and y coordinate norm_x = (x[i] - xmin) / (xmax-xmin) norm_y = (y[i] - ymin) / (ymax-ymin) # check if the point lies in the grid if ( (norm_x >= 0) & (norm_x < 1) & (norm_y >= 0) & (norm_y < 1) ): # calculate the indices in the 2d grid index_x = numpy.int(norm_x * grid_width); index_y = numpy.int(norm_y * grid_height); # apply the operation grid[index_x, index_y] = operation(grid[index_x, index_y], aux[i] if aux is not None else None) # To make a 2d histogram of 10 by 20 cells: # data_x and data_y are 1d numpy arrays containing the data, and # xmin, xmax, ymin, ymax define the border of the grid shape = (10,20) counts = np.zeros(shape) statistic2d(counts, data_x, data_y, None, xmin, xmax, ymin, ymax, operation_count) # To get a 2d grid with a min and max value of data_x at each cell minmax = np.zeros(shape + (2,)) minmax[...,0] = np.inf # Infinity and -infinity are good initial values since they will always be bigger # (or smaller) than any finite value. minmax[...,1] = -np.inf statistic2d(minmax, data_x, data_y, data_x, xmin, xmax, ymin, ymax, operation_minmax) # calculate the standard deviation on a 2d grid for x by calculating the count, the # sum of x and the sum of x**2 at each cell moments012 = np.zeros(shape + (3,)) statistic2d(moments012, data_x, data_y, data_x, xmin, xmax, ymin, ymax, operation_moment_012) # then calculate the raw moments moments2 = moments012[...,2] / moments012[...,0] moments1 = moments012[...,1] / moments012[...,0] # and finally the standard deviation (non stable algorihm) std = numpy.sqrt((moments2 - moments1**2))
All Tables
Functions which can be computed on N dimensional grids and exploit the fast binning algorithm, which are readily available in vaex.
All Figures
 |
Fig. 1. Comparison between scatter and density plots when trying to visualize the sky as observed by Gaia DR1. Top left: a scatter plot showing 10 000 sources in Equatorial coordinates, which does reveal some structure in the disk. Top right: idem, with 1 000 000 sources, hiding almost any structure present in the data. Bottom: a density plot with 1 142 679 769 sources, where the logarithm of the density is colour-mapped (black is low density and white is high density). This reveals much more features in the data, such as structure in the disk and artefacts related to the scanning nature of the Gaia satellite. |
| In the text | |
 |
Fig. 2. Example of one dimensional visualization for the vaex-viz package. Top: histogram of Lz, the angular momentum in the z direction. Because the units are specified in the data file for this column, it will be included in the labelling of the x axis by default. Bottom: similar as above, but showing the mean of the energy E in each bin. |
| In the text | |
 |
Fig. 3. Example of two dimensional visualization for the vaex-viz package. Top left: two dimensional histogram, colour coded by a colour map visualizing the counts in each bin. Top right: similar to the left, but showing the standard deviation in each bin. Middle left: a zoom-in of the E versus Lz space, where a rectangular region is selected using vaex’ selection mechanism. The underlying visualization of all the data is slightly faded out to highlight the selection. Middle right: similar to the top left, but displaying the both the full dataset and the selection, where the selected rectangular region corresponds to a satellite that is not fully phase mixed. Bottom left: similar as the middle right plot but overlaying the mean velocity of the selection using vectors. Bottom right: similar as the bottom left, except now overlaying the velocity dispersion tensor represented by ellipses. |
| In the text | |
 |
Fig. 4. Top: screenshot of a Jupyter notebook using the widget back-end to visualize the dataset interactively using panning, zooming and selection with the mouse. The back-end used here is bqplot. Bottom: a 3d visualization using the ipyvolume backend, showing the stream discussed before, where the vector visualize the mean velocity. An interactive version of this visualization is available online. |
| In the text | |
 |
Fig. 5. Top left: a screenshot of main application window. On the left it shows the open tables, and on the right the meta-data and operations that can be performed on the table. Top right: one dimensional histogram showing the Lz distribution of our example dataset. Bottom left: a two dimensional plotting window, showing E versus Lz. Bottom right: a volume rendering of a stream in our example dataset in Cartesian coordinates. The mean velocity field is also displayed with the help of vectors. |
| In the text | |
 |
Fig. 6. Left: a two dimensional plot window, showing E vs. Lz, similar to the middle left panel of Fig. 3. Right: a two dimensional plot windows, showing x vs. y, similar to the middle right panel of Fig. 5, but sharing the same selection as that shown in the left panel, demonstrating the linked views feature. |
| In the text | |
 |
Fig. 7. Left: ranking of subspaces my mutual information. Right, top row: two subspaces having the highest; bottom row: two subspaces with the lowest mutual information rank. |
| In the text | |
 |
Fig. 8. Examples of density maps created from big datasets. Left: drop-off locations of the Yellow cab taxies in New York City between 2009 and 2015. In total ∼1 billion GPS coordinates are plotted. Middle: density plot of particle position from the Aquarius pure dark matter simulations (Springel et al. 2008, Aquarius A level 2), ∼0.6 billion particles. Right: open street map GPS data, showing ∼2 billion coordinates. |
| In the text | |
 |
Fig. A.1. Python code equivalent of our generalized Nd binning code, but only for 2d for clarity. |
| In the text | |
Current usage metrics show cumulative count of Article Views (full-text article views including HTML views, PDF and ePub downloads, according to the available data) and Abstracts Views on Vision4Press platform.
Data correspond to usage on the plateform after 2015. The current usage metrics is available 48-96 hours after online publication and is updated daily on week days.
Initial download of the metrics may take a while.


