Perhaps the most important recent improvement in echelle reduction
procedures is the introduction of spectral order decomposition. Recall that
we define the spatial profile to be the relative illumination
profile of an order perpendicular to echelle dispersion. For an ideal
optical system that is perfectly aligned, the spatial profile is a
monochromatic image of the illuminated entrance slit, aligned along
detector columns (after re-orientation, if necessary). In this case a
spectral order S can be represented as the product of a spectrum f and
a spatial profile g:
The discrete version of Eq. (3) includes integration
of the spatial profile over each detector pixel:
![$\displaystyle \omega_{x,y}^j = \left\{
\begin{array}[c]{ll}
0 & j<j_0 \\
\text...
...bf{FRACT}\left(y_0(x) M\right) & j=j_0+M \\
0 & j>j_0+M \\
\end{array}\right.$](/articles/aa/full/2002/15/aah3260/img17.gif) |
(5) |
During spectroscopic reduction, our goal is to decompose each
observed order Sx,y into the spectrum fx and a spatial profile
gj. We can accomplish this decomposition by solving the inverse
problem:
Selection of the oversampling factor M needs special attention.
One extreme case occurs when orders are perfectly aligned with detector
rows, in which case no oversampling (M=1) is needed. As the derivative
of y0(x) deviates from zero, M should increase. In order to
decouple the value of M from varying order inclination (thereby making
the problem tractable), we introduce an additional constraint:
(6):
| (9) |
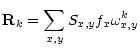 |
(11) |
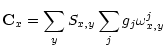 |
(12) |
![$\displaystyle {\bf D}_x = \sum_y \left[ \sum_j g_j \omega_{x,y}^j\right]^2$](/articles/aa/full/2002/15/aah3260/img25.gif) |
(13) |
The very simple structure of
![]() makes it easy to construct
the matrix
makes it easy to construct
the matrix
![]() for every x. This
block-diagonal matrix is the key to efficient computations. Each square
block has M+1 elements on a side and is proportional to the product of
two scalars, such as
for every x. This
block-diagonal matrix is the key to efficient computations. Each square
block has M+1 elements on a side and is proportional to the product of
two scalars, such as
![]() ,
for example.
Therefore
,
for example.
Therefore ![]() does not have to be computed for every value of x.
Only the first M+1 elements are needed! The regular shape of
does not have to be computed for every value of x.
Only the first M+1 elements are needed! The regular shape of
![]() greatly simplifies the evaluation of
matrix
greatly simplifies the evaluation of
matrix
![]() in Eq. (8), making it possible to use fast
and efficient numerical methods to solve for the spatial profile.
in Eq. (8), making it possible to use fast
and efficient numerical methods to solve for the spatial profile.
Iteration is used to solve Eqs. (8)-(13).
Beginning with an initial guess for the spectrum (e.g.
![]() ), we solve Eq. (8) to obtain the spatial profile,
normalize the total area, and then solve Eq. (9) to obtain an
improved estimate of the spectrum. Iteration ceases when the maximum
fractional change in the spectrum becomes small.
), we solve Eq. (8) to obtain the spatial profile,
normalize the total area, and then solve Eq. (9) to obtain an
improved estimate of the spectrum. Iteration ceases when the maximum
fractional change in the spectrum becomes small.
The decomposition procedure produces the best possible spectrum and
slit function, if the order location
![]() is accurately determined
and large scale structure in the image is dominated by the spectrum.
Local defects like cosmic rays, bad pixels, and noise have minimal
impact on the results! This can be illustrated by comparing an observed
order with a model order constructed from the derived f and g,
convolved with detector pixels according to Eq. (4)
(Figs. 4 and 5). The only type of
artifact that adversely affects decomposition are rows of bad pixels
with length and orientation similar to spectral orders. Fortunately,
such defects can usually be identified a priori and ignored during
decomposition, leaving no trace of the bad row (Fig. 4).
is accurately determined
and large scale structure in the image is dominated by the spectrum.
Local defects like cosmic rays, bad pixels, and noise have minimal
impact on the results! This can be illustrated by comparing an observed
order with a model order constructed from the derived f and g,
convolved with detector pixels according to Eq. (4)
(Figs. 4 and 5). The only type of
artifact that adversely affects decomposition are rows of bad pixels
with length and orientation similar to spectral orders. Fortunately,
such defects can usually be identified a priori and ignored during
decomposition, leaving no trace of the bad row (Fig. 4).
Generalization of the decomposition procedure to handle tilted or
curved slit images is straightforward, if the image geometry is known.
The only difference is that the mapping between f and detector pixel
is a function of both x and y, making the pattern of
![]() slightly more complicated. Nevertheless, the structure of
slightly more complicated. Nevertheless, the structure of
![]() remains
block-diagonal. If the PSF has a 2D structure, for example when an image
slicer is used, the order can still be decomposed using a 2D
representation of the PSF in place of the 1D spatial profile. For UVES,
deviations from a 1D representation of g cause errors smaller than the
typical readout noise.
remains
block-diagonal. If the PSF has a 2D structure, for example when an image
slicer is used, the order can still be decomposed using a 2D
representation of the PSF in place of the 1D spatial profile. For UVES,
deviations from a 1D representation of g cause errors smaller than the
typical readout noise.
REDUCE uses the decomposition routine to (a) normalize flat field images and derive order shape functions, (b) locate the inter-order gaps and estimate scattered light, and (c) extract spectral orders.
Copyright ESO 2002